1. Single-cell sequencing
Single-cell sequencing is a technology that provides a snapshot of the transcriptome (i.e., the set of gene expressions) of a collection of individual cells. We present a brief ovewview of single-cell sequencing.
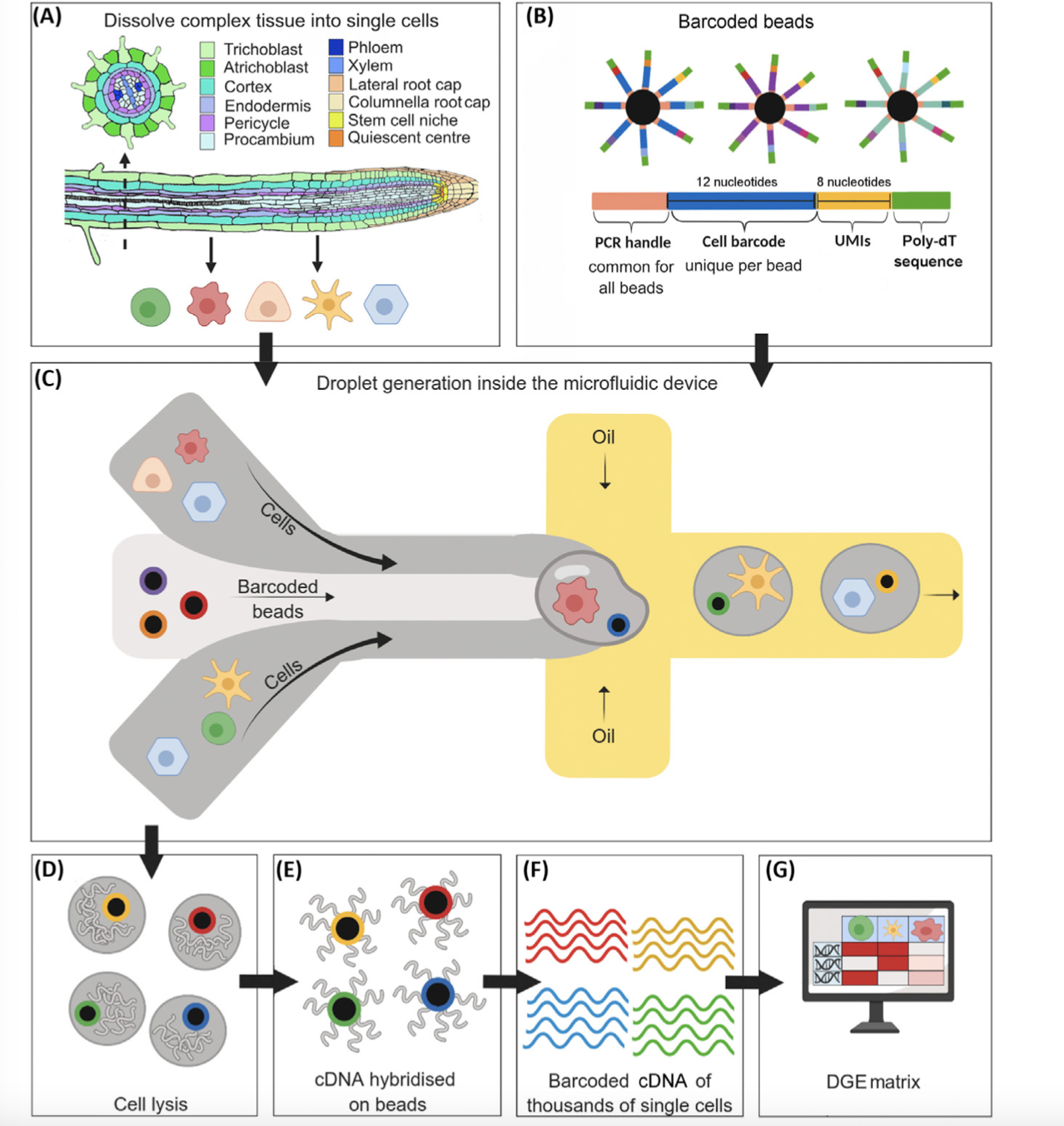
- a) Start with some tissue or collection of cells. Often, the tissue consists of cells of many types, and the goal is to use single-cell sequencing to identify those cell types.
- (b) Generate lots of barcoded beads. Each barcoded bead consists of two parts: (i) a bead, and (ii) a short oligonucleotide (or “oligo”). The oligos are tethered to the bead, like hairs on a person’s head.
The oligo is made of up several smaller parts.
- PCR handle – this is required for the PCR and sequencing steps; it is common across all oligos on all beads.
- Cell barcode – this identifes the cell; importantly, the cell barcode is the same across all oligos on a given bead.
- UMI (unique molecular identifier) – each oligo on a given bead has a distinct UMI. The UMI eliminates bias that arises during the PCR and sequencing steps.
- Poly-dt sequece – a long sequence of T nucleotides. This captures the mRNA transcripts, which are polyadenylated after transcription.
(c) Using a microfluidic device, trap each cell inside a droplet along with a barcoded bead.
(d) Lyse the cells within the droplets.
(e) The mRNAs bind the oligos. Use reverse transcription to generate cDNAs hybridized to the bead surface.
(f) Wash the cDNAs away from the beads, and sequence in bulk (see this post for a review of sequencing).
(g) Use software (e.g., cellranger) to generate the cell-by-gene expression matrix. This roughly involves (i) collapsing all reads with the same UMI into a single read, (ii) determining the cell from which a read came, and (iii) mapping the read onto a reference genome to determine the gene from which the read came.
2. Large-scale single-cell sequencing
There is a major limitation to standard (or small-scale) single-cell sequencing: each droplet can contain at most one cell. If multiple cells are captured in a single droplet, then their transcriptomes are impossible to distinguish.
Experimenters do not have exact control over how many cells are captured in a droplet; this quantity is Poisson-distributed. By experimental design, the majority of droplets contain zero cells.
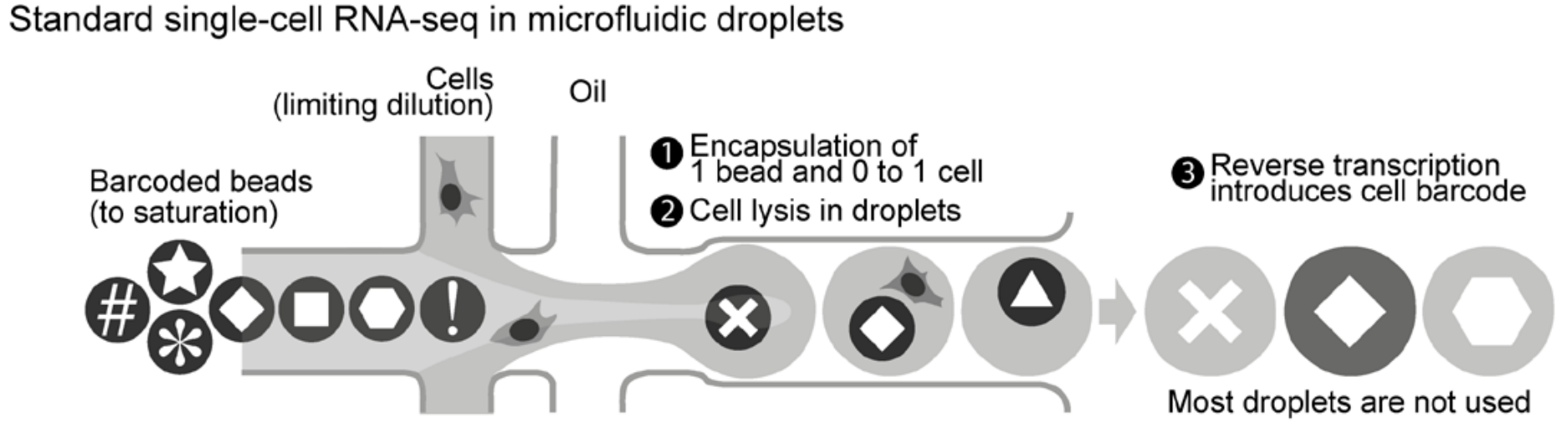
- A new strategy called scifi-RNA-seq rectifies this inefficiency.
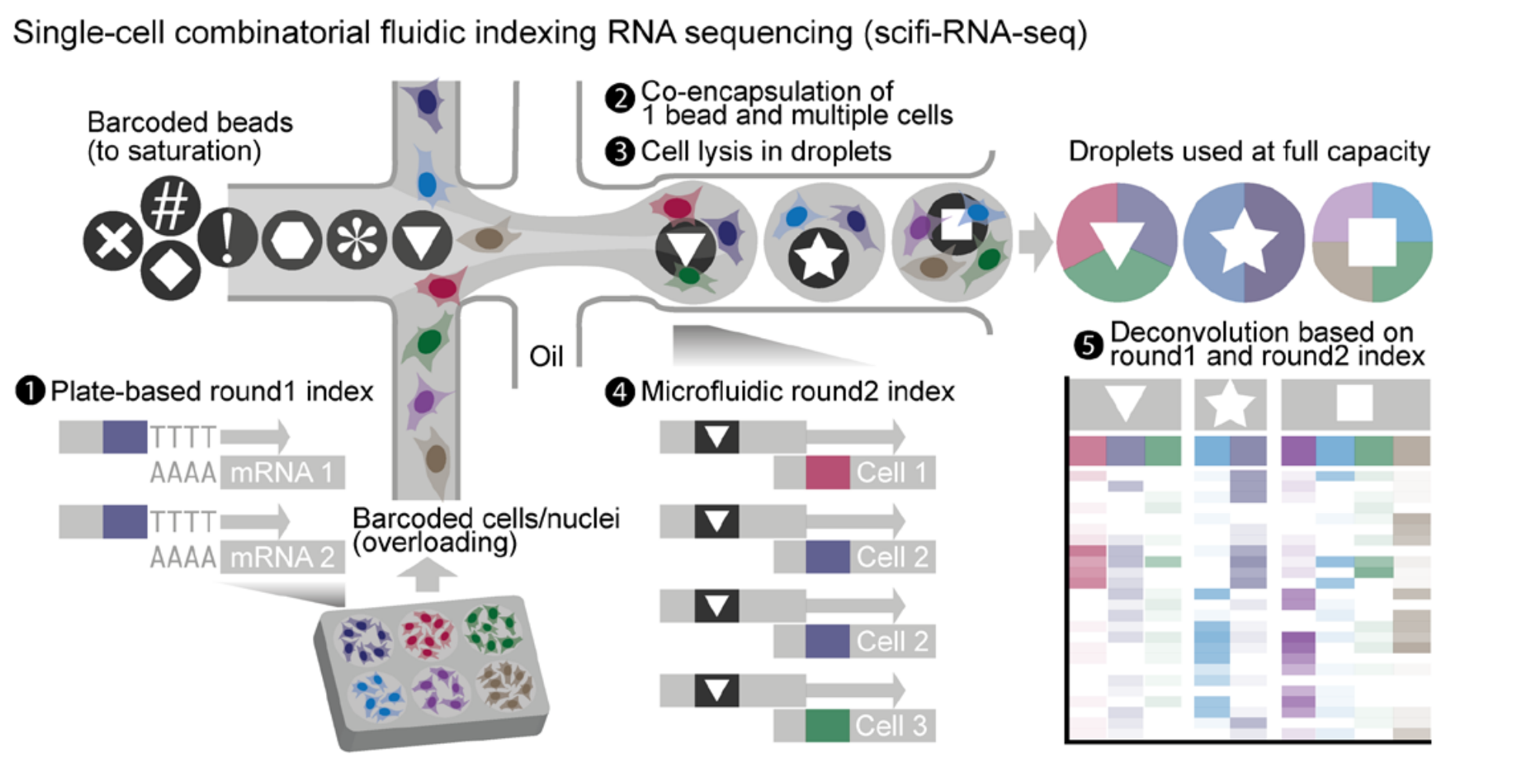
- The cells are divided into groups on a microwell plate (i.e., a plate with a bunch of wells). The membrane of each cell is permeabilized (not lysed), and the entire transcriptome is barcoded with a round-1 barcode and reverse transcribed into cDNA. A UMI and PCR handle are also added during this step. In the figure, the round-1 barcodes are shown in color.
- Using a microfluidic device, capture multiple cells inside a droplet along with a barcoded bead.
- Lyse the cells inside the droplets.
- Ligate a round-2 barcode onto the cDNAs. In the figure, the round-2 barcodes are represented as shapes.
- Pool together the cDNAs and sequence them in bulk. Use software to generate the cell-by-gene expression matrix. Use both the round-1 and round-2 barcodes to determine the cell from which a transcript came.
3. Applications
Large-scale single-cell sequencing enables exciting new applications in single-cell CRISPR screens, cell atlases, and population genomics.
sci-RNA-seq3 (a method related to scifi-RNA-seq) was used to generate an atlas of human fetal tissue.
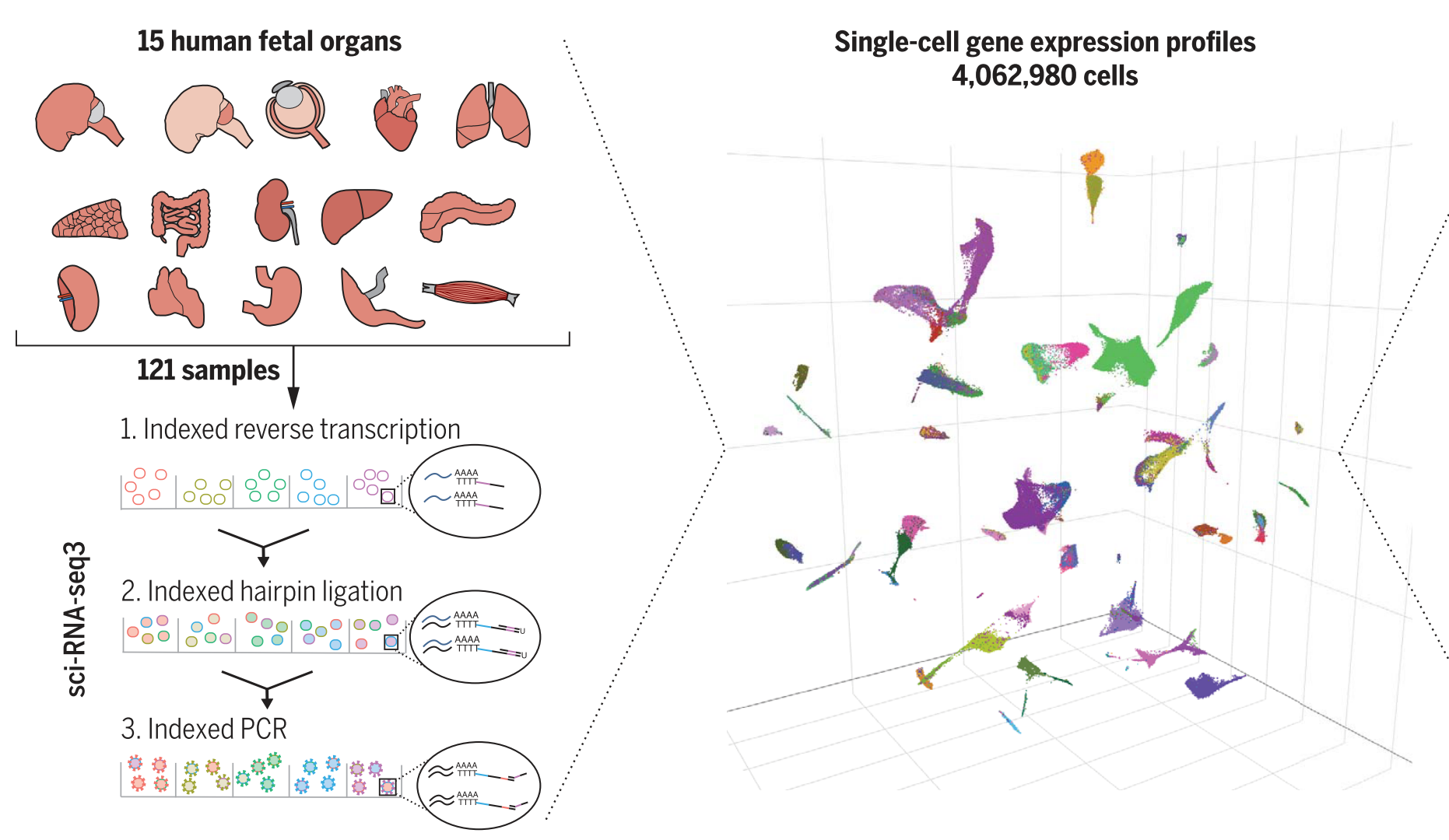
- Large-scale single-cell sequencing opens new opportunities for statisticians, data scientists, and computer scientists.